-Search query
-Search result
Showing 1 - 50 of 1,051 items for (author: xu & th)

EMDB-40180: 
MsbA bound to cerastecin C
Method: single particle / : Chen Y, Klein D

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
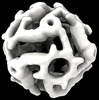
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-41235: 
VMAT1 dimer with MPP+ and reserpine
Method: single particle / : Ye J, Liu B, Li W

EMDB-41236: 
VMAT1 dimer with amphetamine and reserpine
Method: single particle / : Ye J, Liu B, Li W

EMDB-41237: 
VMAT1 dimer with dopamine and reserpine
Method: single particle / : Ye J, Liu B, Li W
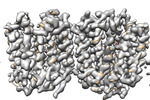
EMDB-41238: 
VMAT1 dimer in unbound form and with reserpine
Method: single particle / : Ye J, Liu B, Li W

EMDB-41239: 
VMAT1 dimer with histamine and reserpine
Method: single particle / : Ye J, Liu B, Li W

EMDB-41240: 
VMAT1 dimer with norepinephrine and reserpine
Method: single particle / : Ye J, Liu B, Li W

EMDB-41241: 
VMAT1 dimer with reserpine
Method: single particle / : Ye J, Liu B, Li W

EMDB-41242: 
VMAT1 dimer with serotonin and reserpine
Method: single particle / : Ye J, Liu B, Li W

PDB-8tgj: 
VMAT1 dimer in unbound form and with reserpine
Method: single particle / : Ye J, Liu B, Li W

PDB-8tgl: 
VMAT1 dimer with norepinephrine and reserpine
Method: single particle / : Ye J, Liu B, Li W

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

EMDB-42906: 
Computational Designed Nanocage O43_129
Method: single particle / : Weidle C, Kibler RD

EMDB-42944: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

PDB-8gel: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

PDB-8tl7: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

PDB-8v3b: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

EMDB-16791: 
Photorhabdus luminescens TcdA1 prepore-to-pore intermediate, K1179W mutant
Method: single particle / : Nganga PN, Roderer D, Belyy A, Prumbaum D, Raunser S

EMDB-16792: 
Photorhabdus luminescens TcdA1 prepore-to-pore intermediate, K567W K2008W mutant
Method: single particle / : Nganga PN, Roderer D, Belyy A, Prumbaum D, Raunser S

EMDB-16793: 
Photorhabdus luminescens TcdA1 prepore-to-pore intermediate, C16S, C20S, C870S, T1279C mutant
Method: single particle / : Nganga PN, Roderer D, Belyy A, Prumbaum D, Raunser S

EMDB-16933: 
Structure of cGAS in complex with SPSB3-ELOBC
Method: single particle / : Xu PB, Ablasser A

EMDB-16936: 
cGAS-Nucleosome in complex with SPSB3-ELOBC (composite structure)
Method: single particle / : Xu PB, Ablasser A

EMDB-16937: 
Consensus refinement of cGAS/spsb3/EloBC/Nucleosome
Method: single particle / : Xu PB, Ablasser A

EMDB-16938: 
human cGAS/spsb3/EloBC in complex with nucleosome (2:2)
Method: single particle / : Xu PB, Ablasser A

PDB-8okx: 
Structure of cGAS in complex with SPSB3-ELOBC
Method: single particle / : Xu PB, Ablasser A

PDB-8ol1: 
cGAS-Nucleosome in complex with SPSB3-ELOBC (composite structure)
Method: single particle / : Xu PB, Ablasser A
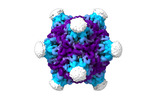
EMDB-19024: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
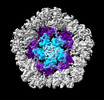
EMDB-19025: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
EMDB-19026: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
EMDB-19027: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb3: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb4: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb5: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb7: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
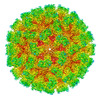

EMDB-43222: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18
Method: single particle / : Sun C, Jiang W

EMDB-43292: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
Method: single particle / : Sun C, Jiang W

EMDB-43293: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18 without DTT treatment
Method: single particle / : Sun C, Jiang W

PDB-8vgr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18
Method: single particle / : Sun C, Jiang W

PDB-8vjr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
Method: single particle / : Sun C, Jiang W
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search










 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
